Research Overview—
Research
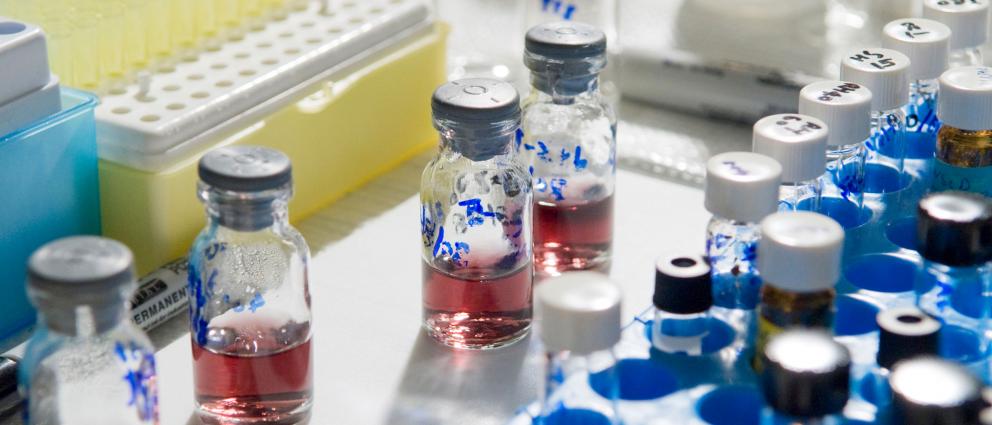
Research in the Wang laboratory focuses on pharmaceutical and exosome analysis, quantitative proteomics, drug metabolism and pharmacokinetics, cancer biomarkers, biotechnology and drug delivery, and drug discovery for parasitic infectious diseases.
1. Blood Exosomal Proteins as Non-Invasive Biomarkers for Liver Cancer and Precision Medicine
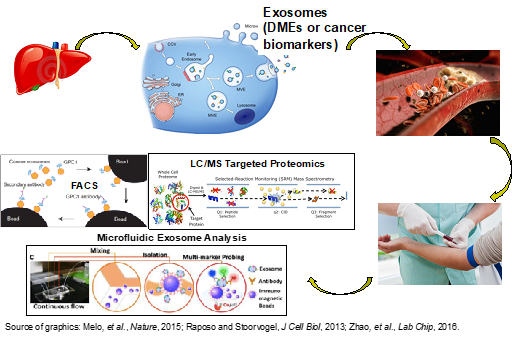
In this project, we aim to identify exosomal proteins that can be used as non-invasive biomarkers (or “liquid biopsy”) for early detection of liver cancer and for drug dose prediction based on hepatic expression of drug-metabolizing enzymes (DMEs). Liquid Chromatography-Mass Spectrometry (LC-MS)-based quantitative proteomic methods are employed to identify exosomal proteins (global proteomics) and achieve robust protein quantification (targeted proteomics). Global and targeted proteomics utilize the state-of-the-art instrumentation, e.g., nanoUPLC-quadrupole time-of-flight (Q-Tof) mass spectrometer, linear ion trap (LTQ) with a Fourier Transform Ion Cyclotron Resonance (FT-ICR) mass spectrometer, and triple quadrupole (QqQ) mass spectrometer. To achieve clinical application, immunoaffinity and microfluidics-based approaches are being developed to isolate liver-specific blood exosomes and afford convenient and ultrasensitive quantification of exosomal biomarkers.
See our research featured on KU in the news: Researchers see exosomes as way to make ‘personalized medicine’ easier and more precise
2. Targeting Cytochrome P450 Enzymes for Leishmaniasis Control
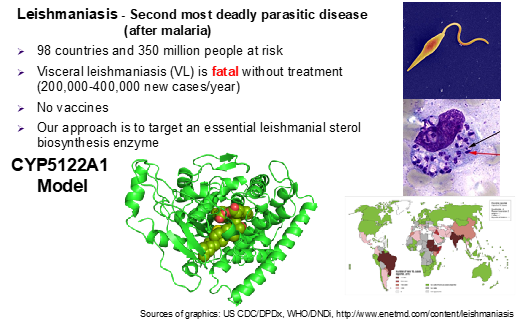
Human leishmaniasis is a devastating infectious disease caused by protozoan parasites belonging to the genus Leishmania. Transmitted by the bite of phlebotomine (bloodsucking) sandflies, these unicellular eukaryotic parasites cause disfiguring skin sores (cutaneous leishmaniasis or CL) and life-threatening infection of vital internal organs (visceral leishmaniasis or VL). VL is the most dangerous and fatal form of the disease, with a mortality rate close to 100% if left untreated. It also is the second most deadly parasitic disease (after malaria) and is responsible for an estimated 0.2 to 0.4 million infections each year worldwide. Leishmaniasis also threatens U.S. military personnel deployed in endemic regions, notably Iraq and Afghanistan. Current antileishmanial drugs have limited efficacy, serious side effects and high cost, and there are no vaccines. Hence, there is a major unmet medical need for safe and effective drugs against leishmaniasis. Our research focuses on understanding biochemical roles of an essential cytochrome P450 (CYP) enzyme (CYP5122A1) in the ergosterol biosynthesis and targeting this enzyme for drug discovery and development for leishmaniasis control
3. Targeted Proteomics to Study Ontogeny of Drug-metabolizing Enzymes, Drug Transporters and Their Regulatory Proteins
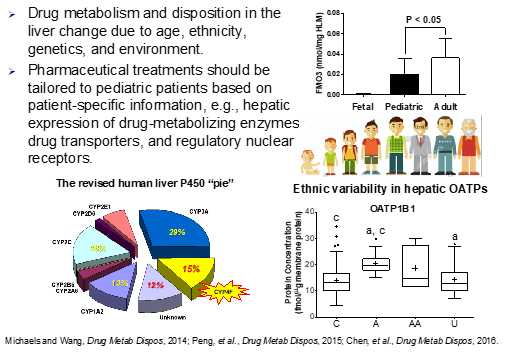
Absolute protein quantification of drug metabolizing enzymes (DMEs), drug transporters (DTs) and their regulatory nuclear receptors (NRs) is important for the characterization of developmental expression patterns and determination of environmental and regulatory influences. Absolute DME and DT quantification also are critical inputs for mechanistic physiologically-based pharmacokinetic (PBPK) models aimed at predicting drug exposure and drug response in diverse populations (e.g., pediatric, geriatric and ethnic groups). Traditionally, absolute protein quantification has relied on antibody-based immunoquantification (i.e., Western blot) using purified proteins as calibration standards. However, Western blot-based immunoquantification has many limitations (e.g., low throughput, cross-reactivity, narrow dynamic range and poor reproducibility). To overcome these limitations, our laboratory was one of the first to develop targeted quantitative proteomic methods for the absolute quantification of CYPs and most recently, DTs, flavin-containing monooxygenases (FMOs) and NRs. We aim to expand the bioanalytical toolbox to gain a better understanding of developmental expression patterns and environmental/regulatory influences on clinically important, as well as under-recognized, DMEs and DTs. Ultimately, this knowledge will inform the rational use of drugs in children of all ages.

